 
Ram usage when running at the first minute is 6656mb, after 8mb and then it out of ram and error. STEP 1 / 1 (thread 0)m/usr/bin/mafft: line 2719: 3192 Killed " $prefix/disttbfast" -q $npickup -E $cycledisttbfast -V "-"$gopdist -s $unalignlevel $legacygapopt $mergearg -W $tuplesize $termgapopt $outnum $addarg $add2ndhalfarg -C $numthreads-$numthreadstb $memopt $weightopt $treeinopt $treeoutopt $distoutopt $seqtype $model -g $gexp -f "-" $gop -Q $spfactor -h $aof $param_fft $algopt $treealg $scoreoutarg $anchoropt pre 2> "$progressfile"ĭoesnt matter I do not enter -thread 4 or not, -maxiterate 2 or -maxiterate 1000, it still shows me same error. doneĬonstructing a UPGMA tree (efffree=1). Generating a scoring matrix for nucleotide (dist=200). mafft_results/Homo_sapiens_1_Pan_troglodytes_1_output.fa Now about my large file: mafft -thread 4 -maxiterate 2 Homo_sapiens_1_Pan_troglodytes_1.fa >. 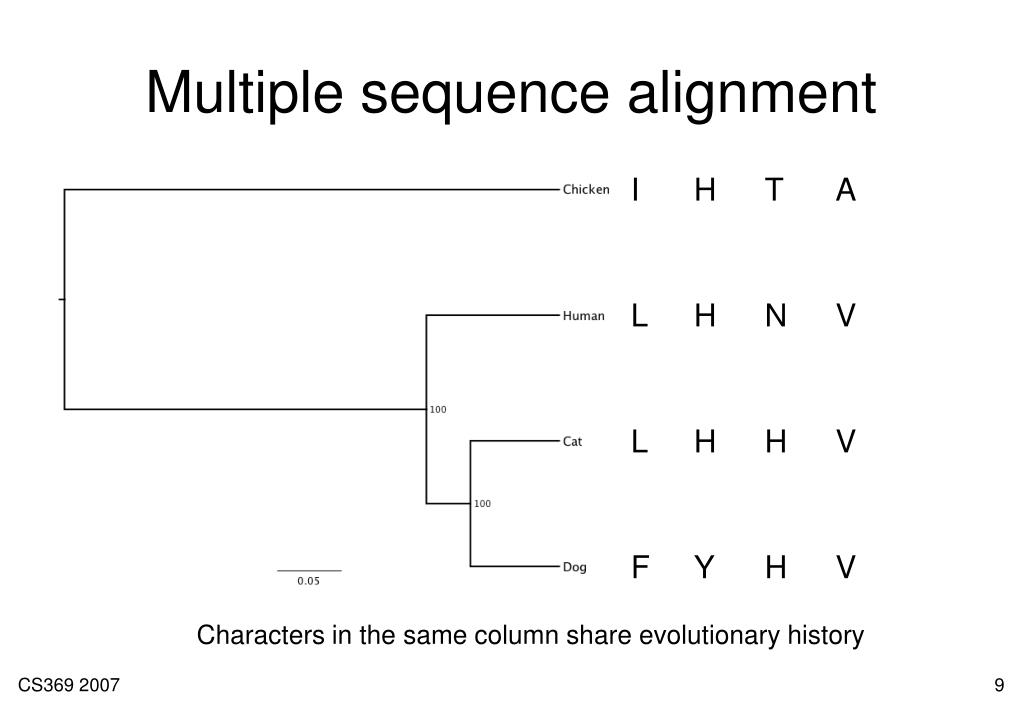
Ttatccactgatcgactcagatcgactcagacatcgatcagctac-gatgac Tacgcatagtagatcgaatcgactagctcagatcgactagatcgatcgt atcgatcgat-gactacgatcgactacgtacgactagctagc MAFFT working cool, when I tried to use it on test sequence file >test1Īnd after MAFFT mafft test.fa > output.fa it returned Why? How could I align it for further comparing? usr/bin/mafft: line 2719: 6030 Killed " $prefix/tbfast" _ -u $unalignlevel $localparam -C $numthreads $seqtype $model -g $pgexp -f $pggop -Q $spfactor -h $pgaof -A $usenaivepairscore $focusarg _ -+ $iterate -W $minimumweight -V "-" $gopdist -s $unalignlevel $legacygapopt $mergearg $outnum $addarg $add2ndhalfarg -C $numthreadstb $rnaopt $weightopt $treeinopt $treeoutopt $distoutopt $seqtype $model -f "-"$gop -Q $spfactor -h $aof $param_fft $localparam $algopt $treealg $scoreoutarg $focusarg /dev/null 2> "$progressfile" mafft_results/Homo_sapiens_1_Pan_troglodytes_1_output.faĪnd it returns me generating a scoring matrix for nucleotide (dist=200). mafft -globalpair -maxiterate 1000 Homo_sapiens_1_Pan_troglodytes_1.fa >. 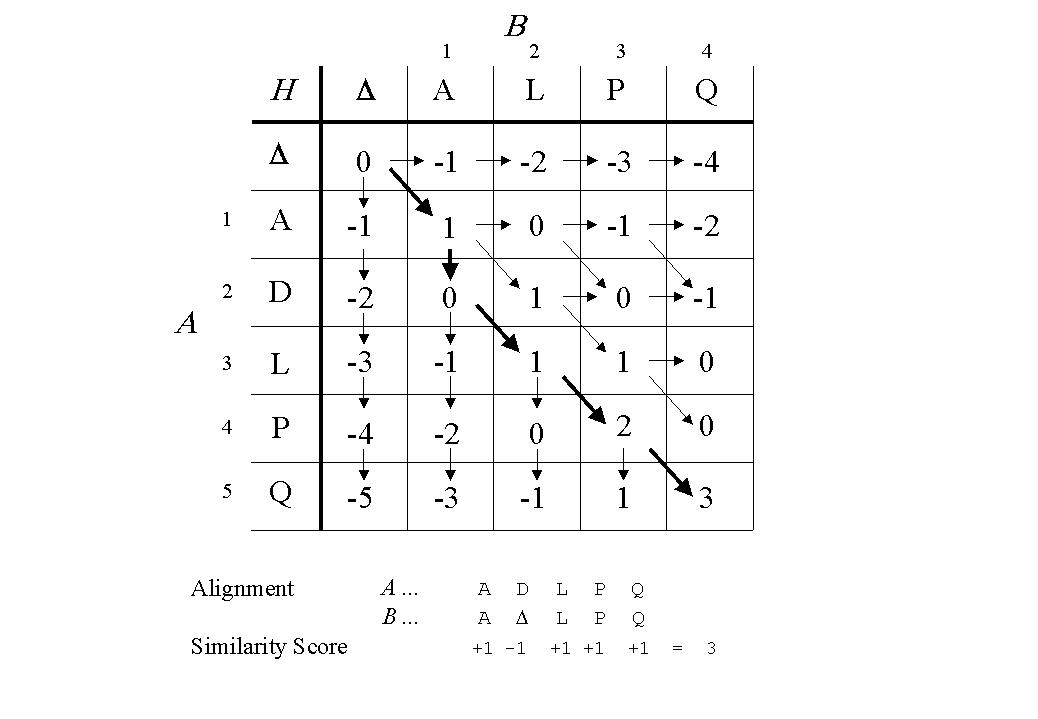
This file is 461mb (1 chromosome of homo sapiens & 1 chromosome of pan troglodytes) and I want to align it. fa sequenes like: >some description of first.ĪCTGACTACGTACGACTACGATCTGACTACACGTAGATAGACTAGTCACTACG 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
